|
CEPS
24.01
Cardiac ElectroPhysiology Simulator
|
|
CEPS
24.01
Cardiac ElectroPhysiology Simulator
|
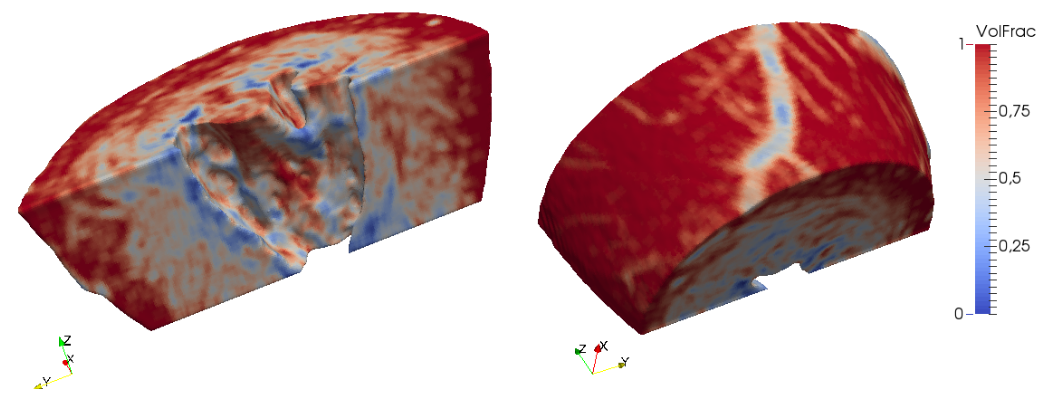
Volume fraction is a way to include cell-scale heterogeneities of the myocardium into tissue scale computations.
Citing [1]:
"The standard model used in cardiac electrophysiology is the bidomain model. It is an averaged model derived from the microscopic properties of the tissue. The bidomain model assumes that the electrically active myocytes are present uniformly everywhere in the heart. While this is a reasonable assumption for healthy hearts, it fails in some pathological cases where significant changes in the tissue structure occur, for example in ischaemic and rheumatic heart disease, inflammation, hypertrophy, or infarction. These tissue heterogeneities are often taken into account through an ad-hoc tuning of model parameters. [...]
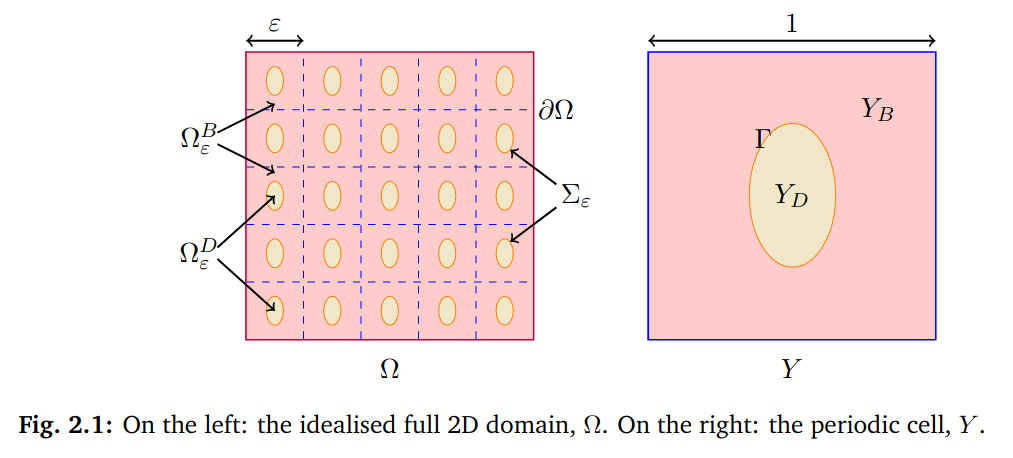
We assume[...] a periodic alternation of healthy (bidomain model) and altered (diffusive inclusion) tissue patches. Such a model may be simulated directly, at the high computational cost of a very fine discretisation. Instead we derive[...] a homogenized model at the macroscopic scale, using a rigorous two-scale analysis. We recover[...] a bidomain-type model with modified conductivity coefficients. "
Volume fraction (VF) is the ratio between normal tissue and diffusive inclusions.
In order to add volume fraction in your computations (e.g. for scarred tissue), at least two data sources must be provided:
Each file is made of 10 columns: the volume fraction, then the modified values of conductivity tensors
Here is an example of interpolation map file
[1] Andjela Davidović. Multiscale mathematical modelling of structural heterogeneities in cardiac electrophysiology. PhD Thesis. Université de Bordeaux, 2016.